Blocking quorum sensing would not in theory kill the bacteria, but it would give more time for the bodies immune system to recognize and destroy the bacteria. This is starting to be a more common approach in the design of anti-microbials, and in virus vaccines. A slower and less virulent form of both virus's and bacteria can sometimes be dealt with easily enough by the body's own immune system as it has more time to respond to the invading challenge.
There is also an evolutionary reason for the effectiveness of quorum sensing disruption over conventional antibiotics. Antibiotics kill off any bacteria that aren't resistant, creating a highly competitive growth environment with a large selection bias against non-resistant bacteria. Disrupting the quorum sensing system on the other hand should mean that the bacteria are still growing in a non-selecting environment, right up to the point where the body's defense system kills them off. There is less evolutionary pressure on them to develop resistance.
A paper in PLoS (reference below) looked at developing a method for how (and whether) resistance towards quorum sensing disruption might occur. They chose to base their model on the effects of altering the genes involved in quorum sensing, rather than other methods for quorum sensing disruption, such as blocking or destroying the little molecules used for signalling and one of the first things they found was that there are a large number of variations in these genes, despite the fact that many of the actual signalling molecules are fairly similar.
Lots of gene variation means bad news for gene-based disruption as there is potential for the bacteria to share these genes among themselves, and thus potentially end up with a combination that renders the gene-blocking anti-bacterial invalid. And swopping genes is one of the things bacteria do best.

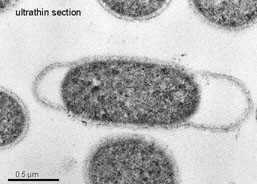
Figure above shows two bacteria connecting together via a long thread-like molecule called a 'pilus'. The genes do not actually travel through the pilus, but the pilus pulls both bacteria close together allowing the gene (in the form of a plasmid) to be passed between them.
Another thing that was examined was the effect of quorum sensing disruption on fitness. The idea that resistance to quorum sensing (which will henceforth be called QS as I can't be bothered to keep typing it out) will take more time to develop is based heavily on the idea that tampering with the QS does not affect the general health of the bacteria. Sure it prevents bacteria from communicating, but each individual bacteria is no less likely to thrive, and therefore there is less selection pressure.
However almost all of these experiments (certainly those in academic labs) have been done in conditions of nice safe uber-rich bacterial growth media. Even slightly shaken bacteria may thrive fine on nutrient rich agar. Inside an actual living organism though (the normal living place of pathogenic bacteria) disrupting the QS system has major implications. Not just in the fact that non QS-able organisms are often outcompeted by other bacteria, but also because some QS-controlled proteins seem (under certain conditions) to be vital for bacterial growth. If disruption of QS is leading to bacterial death then this puts a massive evolutionary pressure on bacteria to evolve resistance, similar to the pressure to evolve antibiotic resistance.
Interestingly (if slightly off topic) it is vaguely suggested that the necessity of QS for growth might help to prevent 'cheaters' within a bacterial population; bacteria that take up all the benefits of it's QS-ing neighbors (i.e biofilm formation, increased virulence etc) without having to expend energy into the QS system.
In view of this data it looks as if, despite it's undoubtable advantages as an antimicrobial, QS-disrupting systems will inevitably succumb to bacterial resistance. In the presence of a selective pressure favoring it resistance will always develop. There is no golden bullet for antibacterial strategies, just a lot of little silver bullets that eventually loose their power.
So as not to end this on a depressing note, it is worth mentioning that the fact that there is a lot of research going into alternative (i.e not antibiotics) antimicrobials is encouraging. Phage therapy and QS disruption will probably never replace antibiotic therapy, but it is worth having as many different strategies as possible to deal with the threat of bacterial infection.
---
Defoirdt T, Boon N, & Bossier P (2010). Can bacteria evolve resistance to quorum sensing disruption? PLoS pathogens, 6 (7) PMID: 20628566
---
Follow me on Twitter!